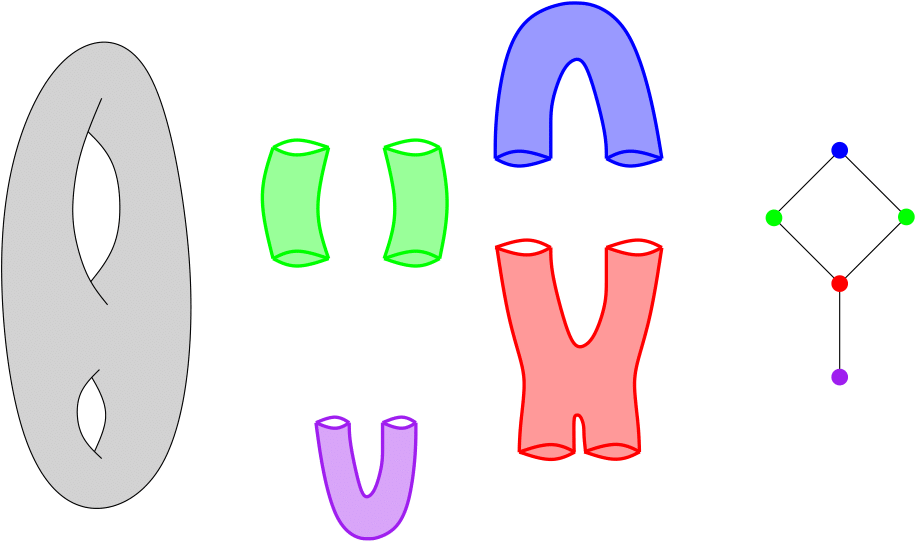- Author
- Mathieu Carrière
Visualizations of the simplicial complexes can be done with either neato (from graphviz), geomview, KeplerMapper. Input point clouds are assumed to be OFF files
Covers
Nerves and Graph Induced Complexes require a cover C of the input point cloud P, that is a set of subsets of P whose union is P itself. Very often, this cover is obtained from the preimage of a family of intervals covering the image of some scalar-valued function f defined on P. This family is parameterized by its resolution, which can be either the number or the length of the intervals, and its gain, which is the overlap percentage between consecutive intervals (ordered by their first values).
Nerves
Nerve definition
Assume you are given a cover C of your point cloud P. Then, the Nerve of this cover is the simplicial complex that has one k-simplex per k-fold intersection of cover elements. See also Wikipedia .
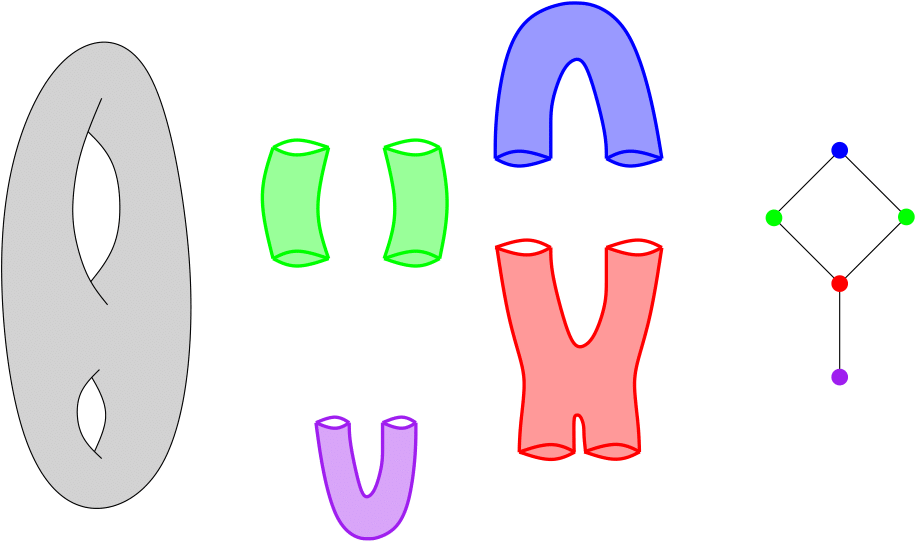
Nerve of a double torus
Example
This example builds the Nerve of a point cloud sampled on a 3D human shape (human.off). The cover C comes from the preimages of intervals (10 intervals with gain 0.3) covering the height function (coordinate 2), which are then refined into their connected components using the triangulation of the .OFF file.
#include <gudhi/GIC.h>
#include <string>
#include <vector>
void usage(int nbArgs, char *const progName) {
std::cerr << "Error: Number of arguments (" << nbArgs << ") is not correct\n";
std::cerr << "Usage: " << progName << " filename.off coordinate resolution gain [-v] \n";
std::cerr << " i.e.: " << progName << " ../../data/points/human.off 2 10 0.3 -v \n";
exit(-1);
}
int main(int argc, char **argv) {
if ((argc != 5) && (argc != 6)) usage(argc, argv[0]);
using Point = std::vector<float>;
std::string off_file_name(argv[1]);
int coord = atoi(argv[2]);
int resolution = atoi(argv[3]);
double gain = atof(argv[4]);
bool verb = 0;
if (argc == 6) verb = 1;
if (!check) {
std::clog << "Incorrect OFF file." << std::endl;
} else {
if (verb) {
std::clog <<
"Nerve is of dimension " << stree.dimension() <<
" - " << stree.
num_simplices() <<
" simplices - "
std::clog << "Iterator on Nerve simplices" << std::endl;
std::clog << vertex << " ";
}
std::clog << std::endl;
}
}
}
return 0;
}
Simplex Tree data structure for representing simplicial complexes.
Definition: Simplex_tree.h:95
Filtration_simplex_range const & filtration_simplex_range(Indexing_tag=Indexing_tag())
Returns a range over the simplices of the simplicial complex, in the order of the filtration.
Definition: Simplex_tree.h:338
Simplex_vertex_range simplex_vertex_range(Simplex_handle sh) const
Returns a range over the vertices of a simplex.
Definition: Simplex_tree.h:349
size_t num_vertices() const
Returns the number of vertices in the complex.
Definition: Simplex_tree.h:651
size_t num_simplices()
Returns the number of simplices in the simplex_tree.
Definition: Simplex_tree.h:664
Cover complex data structure.
Definition: GIC.h:99
void set_resolution_with_interval_number(int reso)
Sets a number of intervals from a value stored in memory.
Definition: GIC.h:591
Persistence_diagram compute_PD()
Computes the extended persistence diagram of the complex.
Definition: GIC.h:1173
void create_complex(SimplicialComplex &complex)
Creates the simplicial complex.
Definition: GIC.h:1333
void set_type(const std::string &t)
Specifies whether the type of the output simplicial complex.
Definition: GIC.h:188
void find_simplices()
Computes the simplices of the simplicial complex.
Definition: GIC.h:1347
void set_cover_from_function()
Creates a cover C from the preimages of the function f.
Definition: GIC.h:603
void write_info()
Creates a .txt file called SC.txt describing the 1-skeleton, which can then be plotted with e....
Definition: GIC.h:1086
void set_gain(double g=0.3)
Sets a gain from a value stored in memory (default value 0.3).
Definition: GIC.h:597
void set_function_from_coordinate(int k)
Creates the function f from the k-th coordinate of the point cloud P.
Definition: GIC.h:506
void set_color_from_coordinate(int k=0)
Computes the function used to color the nodes of the simplicial complex from the k-th coordinate.
Definition: GIC.h:1009
void set_graph_from_OFF()
Creates a graph G from the triangulation given by the input .OFF file.
Definition: GIC.h:344
bool read_point_cloud(const std::string &off_file_name)
Reads and stores the input point cloud from .(n)OFF file.
Definition: GIC.h:244
void set_verbose(bool verb=false)
Specifies whether the program should display information or not.
Definition: GIC.h:196
When launching:
$> ./Nerve ../../data/points/human.off 2 10 0.3 -v
the program output is:
Min function value = -0.979672 and Max function value = 0.816414
Interval 0 = [-0.979672, -0.761576]
Interval 1 = [-0.838551, -0.581967]
Interval 2 = [-0.658942, -0.402359]
Interval 3 = [-0.479334, -0.22275]
Interval 4 = [-0.299725, -0.0431415]
Interval 5 = [-0.120117, 0.136467]
Interval 6 = [0.059492, 0.316076]
Interval 7 = [0.239101, 0.495684]
Interval 8 = [0.418709, 0.675293]
Interval 9 = [0.598318, 0.816414]
Computing preimages...
Computing connected components...
.txt generated. It can be visualized with e.g. python KeplerMapperVisuFromTxtFile.py and firefox.
5 interval(s) in dimension 0:
[-0.909111, 0.00817529]
[-0.171433, 0.367392]
[-0.171433, 0.367392]
[-0.909111, 0.745853]
0 interval(s) in dimension 1:
Nerve is of dimension 1 - 41 simplices - 21 vertices.
Iterator on Nerve simplices
1
0
4
4 0
2
2 1
8
8 2
5
5 4
9
9 8
13
13 5
14
14 9
19
19 13
25
32
20
32 20
33
33 25
26
26 14
26 19
42
42 26
34
34 33
27
27 20
35
35 27
35 34
42 35
44
44 35
54
54 44
The program also writes a file ../../data/points/human_sc.txt. The first three lines in this file are the location of the input point cloud and the function used to compute the cover. The fourth line contains the number of vertices nv and edges ne of the Nerve. The next nv lines represent the vertices. Each line contains the vertex ID, the number of data points it contains, and their average color function value. Finally, the next ne lines represent the edges, characterized by the ID of their vertices.
Using KeplerMapper, one can obtain the following visualization:

Visualization with KeplerMapper
Graph Induced Complexes (GIC)
GIC definition
Again, assume you are given a cover C of your point cloud P. Moreover, assume you are also given a graph G built on top of P. Then, for any clique in G whose nodes all belong to different elements of C, the GIC includes a corresponding simplex, whose dimension is the number of nodes in the clique minus one. See [25] for more details.
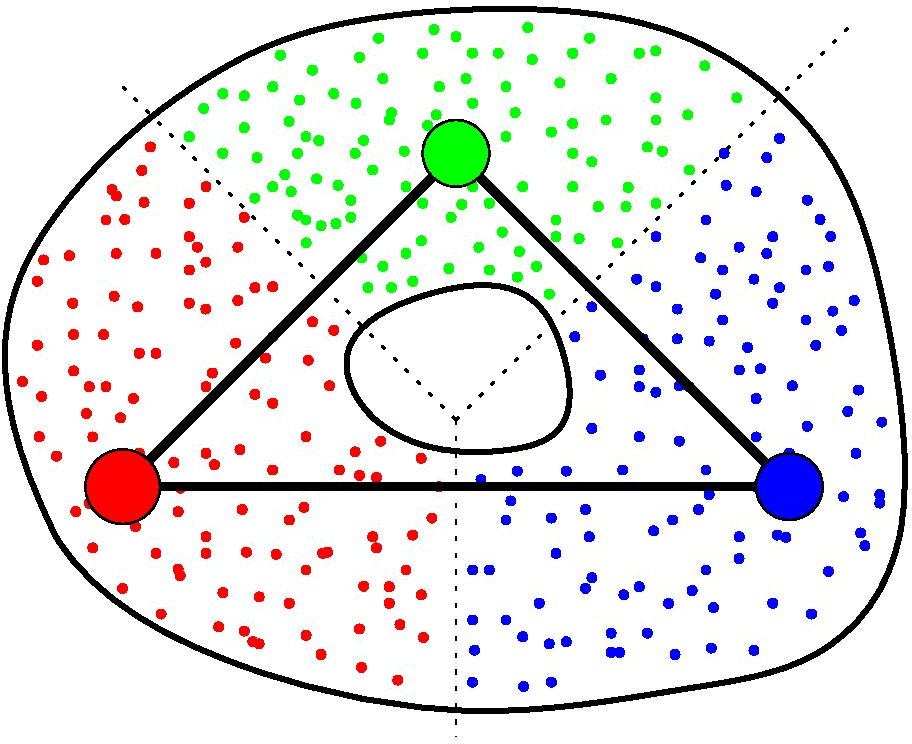
GIC of a point cloud.
Example with cover from Voronoï
This example builds the GIC of a point cloud sampled on a 3D human shape (human.off). We randomly subsampled 100 points in the point cloud, which act as seeds of a geodesic Voronoï diagram. Each cell of the diagram is then an element of C. The graph G (used to compute both the geodesics for Voronoï and the GIC) comes from the triangulation of the human shape. Note that the resulting simplicial complex is in dimension 3 in this example.
#include <gudhi/GIC.h>
#include <string>
#include <vector>
void usage(int nbArgs, char *const progName) {
std::cerr << "Error: Number of arguments (" << nbArgs << ") is not correct\n";
std::cerr << "Usage: " << progName << " filename.off N [-v] \n";
std::cerr << " i.e.: " << progName << " ../../data/points/human.off 100 -v \n";
exit(-1);
}
int main(int argc, char **argv) {
if ((argc != 3) && (argc != 4)) usage(argc, argv[0]);
using Point = std::vector<float>;
std::string off_file_name(argv[1]);
int m = atoi(argv[2]);
bool verb = 0;
if (argc == 4) verb = 1;
if (!check) {
std::clog << "Incorrect OFF file." << std::endl;
} else {
if (verb) {
std::clog <<
"Graph induced complex is of dimension " << stree.dimension() <<
" - " << stree.
num_simplices()
<<
" simplices - " << stree.
num_vertices() <<
" vertices." << std::endl;
std::clog << "Iterator on graph induced complex simplices" << std::endl;
std::clog << vertex << " ";
}
std::clog << std::endl;
}
}
}
return 0;
}
Compute the Euclidean distance between two Points given by a range of coordinates....
Definition: distance_functions.h:32
void set_cover_from_Voronoi(Distance distance, int m=100)
Creates the cover C from the Voronoï cells of a subsampling of the point cloud.
Definition: GIC.h:897
void plot_OFF()
Creates a .off file called SC.off for 3D visualization, which contains the 2-skeleton of the GIC....
Definition: GIC.h:1125
When launching:
$> ./VoronoiGIC ../../data/points/human.off 700 -v
the program outputs SC.off. Using e.g.
$> geomview ../../data/points/human_sc.off
one can obtain the following visualization:
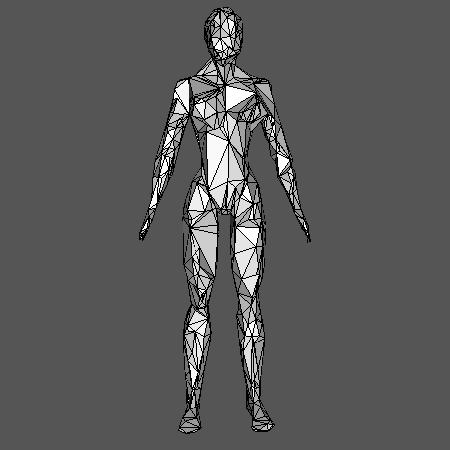
Visualization with Geomview
Functional GIC
If one restricts to the cliques in G whose nodes all belong to preimages of consecutive intervals (assuming the cover of the height function is minimal, i.e. no more than two intervals can intersect at a time), the GIC is of dimension one, i.e. a graph. We call this graph the functional GIC. See [14] for more details.
Example
Functional GIC comes with automatic selection of the Rips threshold, the resolution and the gain of the function cover. See [17] for more details. In this example, we compute the functional GIC of a Klein bottle embedded in R^5, where the graph G comes from a Rips complex with automatic threshold, and the cover C comes from the preimages of intervals covering the first coordinate, with automatic resolution and gain. Note that automatic threshold, resolution and gain can be computed as well for the Nerve.
#include <gudhi/GIC.h>
#include <string>
#include <vector>
void usage(int nbArgs, char *const progName) {
std::cerr << "Error: Number of arguments (" << nbArgs << ") is not correct\n";
std::cerr << "Usage: " << progName << " filename.off coordinate [-v] \n";
std::cerr << " i.e.: " << progName << " ../../data/points/human.off 2 -v \n";
exit(-1);
}
int main(int argc, char **argv) {
if ((argc != 3) && (argc != 4)) usage(argc, argv[0]);
using Point = std::vector<float>;
std::string off_file_name(argv[1]);
int coord = atoi(argv[2]);
bool verb = 0;
if (argc == 4) verb = 1;
if (!check) {
std::clog << "Incorrect OFF file." << std::endl;
} else {
if (verb) {
std::clog <<
"Coordinate GIC is of dimension " << stree.dimension() <<
" - " << stree.
num_simplices()
<<
" simplices - " << stree.
num_vertices() <<
" vertices." << std::endl;
std::clog << "Iterator on coordinate GIC simplices" << std::endl;
std::clog << vertex << " ";
}
std::clog << std::endl;
}
}
}
return 0;
}
double set_automatic_resolution()
Computes the optimal length of intervals (i.e. the smallest interval length avoiding discretization a...
Definition: GIC.h:544
void plot_DOT()
Creates a .dot file called SC.dot for neato (part of the graphviz package) once the simplicial comple...
Definition: GIC.h:1038
double set_graph_from_automatic_rips(Distance distance, int N=100)
Creates a graph G from a Rips complex whose threshold value is automatically tuned with subsampling—s...
Definition: GIC.h:431
void compute_distribution(unsigned int N=100)
Computes bootstrapped distances distribution.
Definition: GIC.h:1236
double compute_p_value()
Computes the p-value, i.e. the opposite of the confidence level of the largest bottleneck distance pr...
Definition: GIC.h:1314
When launching:
$> ./CoordGIC ../../data/points/KleinBottle5D.off 0 -v
the program outputs SC.dot. Using e.g.
$> neato SC.dot -Tpdf -o SC.pdf
one can obtain the following visualization:
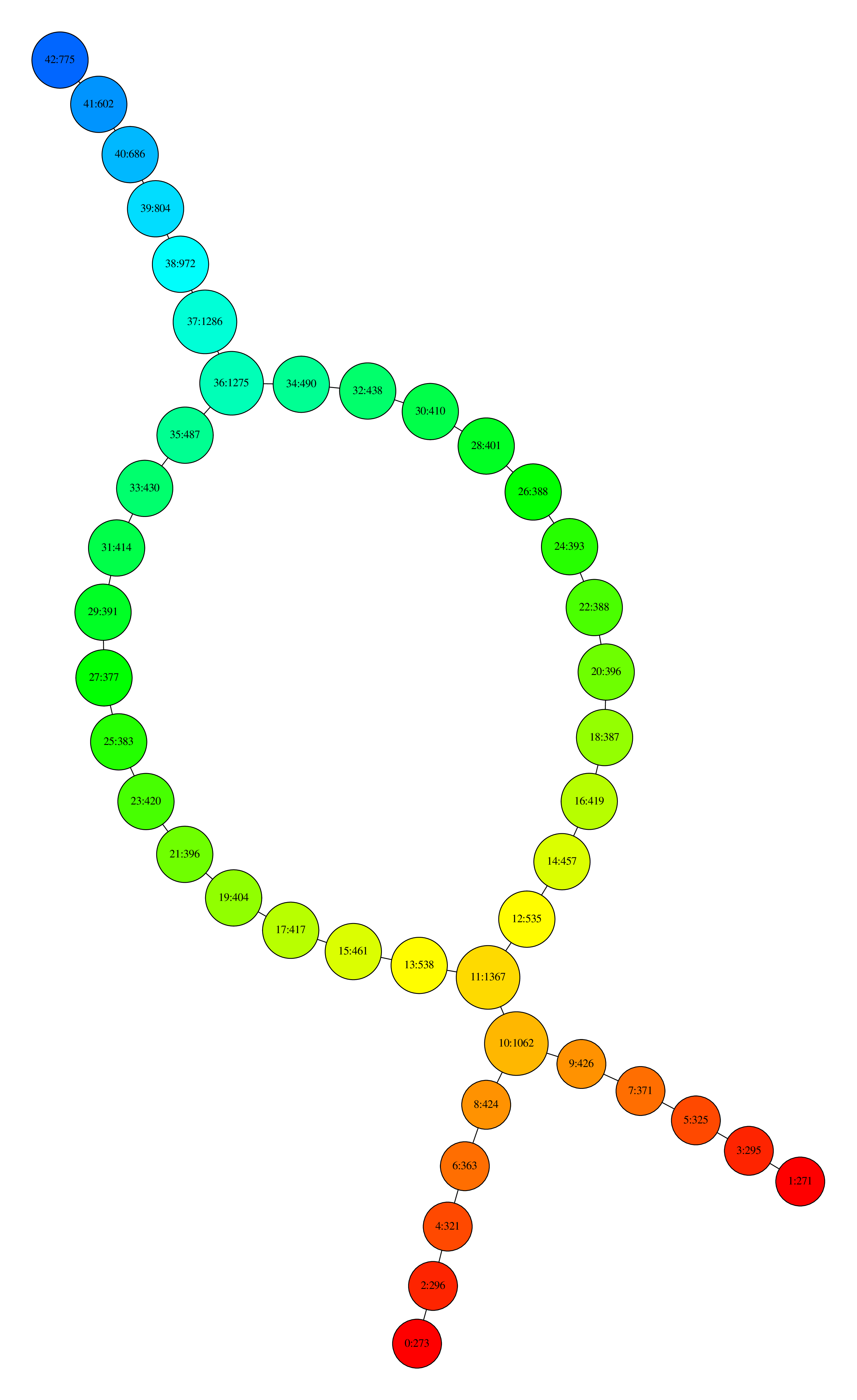
Visualization with Neato
where nodes are colored by the filter function values and, for each node, the first number is its ID and the second is the number of data points that its contain.
We also provide an example on a set of 72 pictures taken around the same object (lucky_cat.off). The function is now the first eigenfunction given by PCA, whose values are written in a file (lucky_cat_PCA1). Threshold, resolution and gain are automatically selected as before.
#include <gudhi/GIC.h>
#include <string>
#include <vector>
void usage(int nbArgs, char *const progName) {
std::cerr << "Error: Number of arguments (" << nbArgs << ") is not correct\n";
std::cerr << "Usage: " << progName << " filename.off function [-v] \n";
std::cerr << " i.e.: " << progName << " ../../data/points/COIL_database/lucky_cat.off "
"../../data/points/COIL_database/lucky_cat_PCA1 -v \n";
exit(-1);
}
int main(int argc, char **argv) {
if ((argc != 3) && (argc != 4)) usage(argc, argv[0]);
using Point = std::vector<float>;
std::string off_file_name(argv[1]);
std::string func_file_name = argv[2];
bool verb = 0;
if (argc == 4) verb = 1;
if (!check) {
std::clog << "Incorrect OFF file." << std::endl;
} else {
if (verb) {
std::clog <<
"Functional GIC is of dimension " << stree.dimension() <<
" - " << stree.
num_simplices()
<<
" simplices - " << stree.
num_vertices() <<
" vertices." << std::endl;
std::clog << "Iterator on functional GIC simplices" << std::endl;
std::clog << vertex << " ";
}
std::clog << std::endl;
}
}
}
return 0;
}
void set_function_from_file(const std::string &func_file_name)
Creates the function f from a file containing the function values.
Definition: GIC.h:485
void set_color_from_file(const std::string &color_file_name)
Computes the function used to color the nodes of the simplicial complex from a file containing the fu...
Definition: GIC.h:989
When launching:
$> ./FuncGIC ../../data/points/COIL_database/lucky_cat.off ../../data/points/COIL_database/lucky_cat_PCA1 -v
the program outputs again SC.dot which gives the following visualization after using neato:
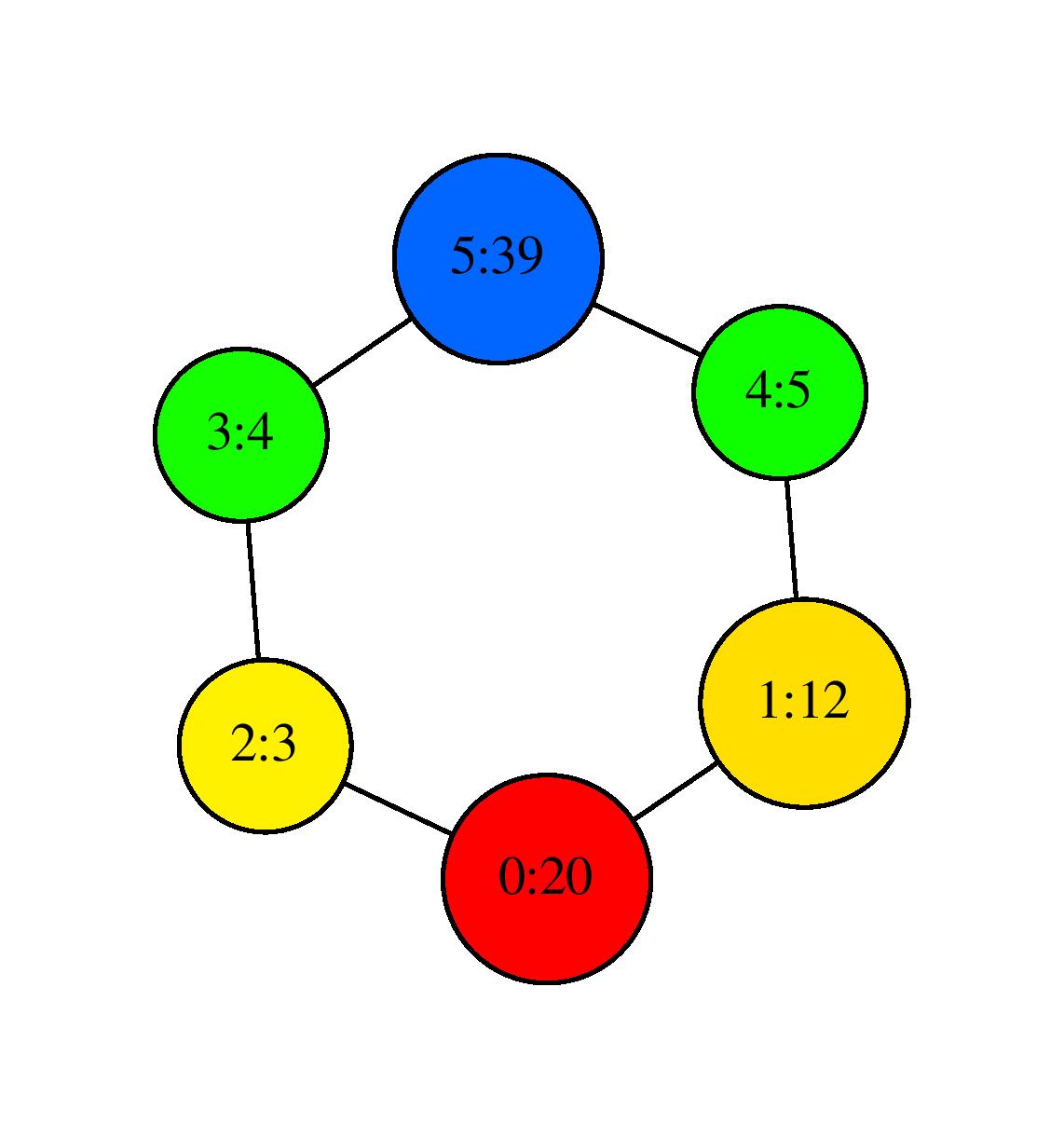
Visualization with neato